|
Gene Ontology) or experimental data (e.g. * Gene-sets, grouping genes on the basis of a-priori knowledge (e.g. * A (ranked) gene list, from a genomic experiment Gene-set enrichment is a data analysis technique taking as input: Nodes represent gene-sets and edges represent mutual overlap in this way, highly redundant gene-sets are grouped together as clusters, dramatically improving the capability to navigate and interpret enrichment results. It will operate on any generic enrichment results as well as specifically on Gene Set Enrichment Analysis (GSEA) results. The EnrichmentMap Cytoscape App allows you to visualize the results of gene-set enrichment as a network. Use the CyRest call to access the aMatReader functionality.*This App is part of the. #scale the data if required RNASeq_expression <- scale(NASeq_expression, center = TRUE, scale = TRUE) #rownames(RNASeq_expression) <- RNASeq_expression_matrix$Name RNAseq_correlation_matrix <- cor( t(RNASeq_expression), method= "pearson") #set the diagonal of matrix to zero - eliminate self-correlation RNAseq_correlation_matrix <- 0 # set all correlations that are less than 0.7 to zero RNAseq_correlation_matrix <- 0 #get rid of rows and columns that have no correlations with the above thresholds RNAseq_correlation_matrix <- RNAseq_correlation_matrix #correlation_filename <- file.path(getwd(), "Data", "Protein_correlation_matrix.txt") write.table( RNAseq_correlation_matrix, file = correlation_filename, col.names = TRUE, row.names = FALSE, sep = " \t ", quote= FALSE) To import our expression data we will match our data-set to the “display name” node attribute. The column “display name” contains gene names which are also found in our Ageing Expression data-set. # "tissue::thyroid gland" "tissue::urine" # "tissue::nervous system" "tissue::pancreas" # "tissue::adrenal gland" "tissue::blood" # "target::development level" "target::family" # "stringdb::interactor score" "stringdb::structures"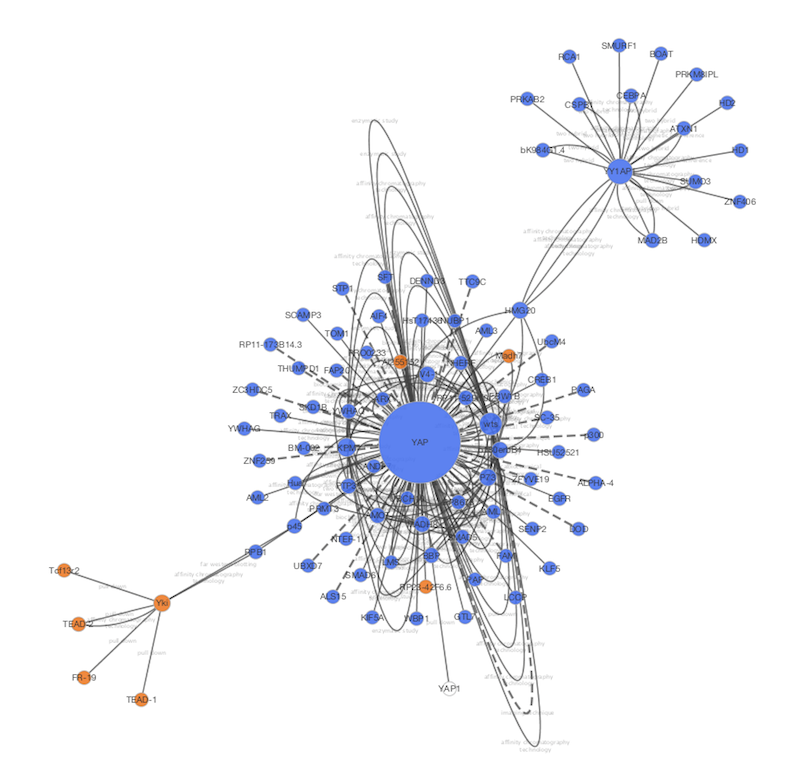  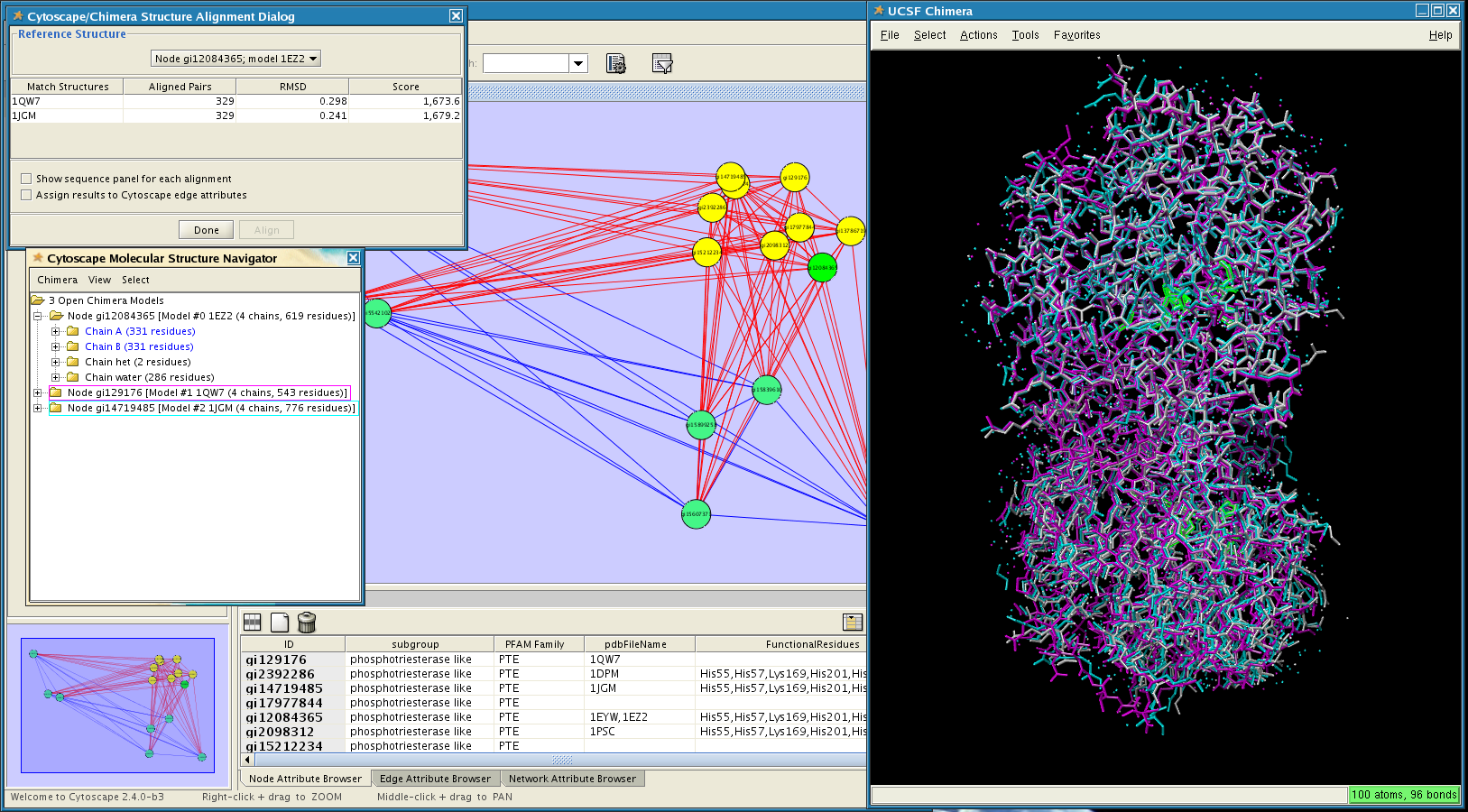
# "compartment::peroxisome" "compartment::plasma membrane" # "compartment::mitochondrion" "compartment::nucleus" # "compartment::golgi apparatus" "compartment::lysosome" # "compartment::endosome" "compartment::extracellular" # "compartment::cytosol" "compartment::endoplasmic reticulum" # "stringdb::enhancedLabel Passthrough" "compartment::cytoskeleton" # "stringdb::species" "stringdb::STRING style" # "stringdb::description" "stringdb::namespace" "stringdb::node type" # "stringdb::full name" "stringdb::database identifier" # "stringdb::canonical name" "display name" GetTableColumnNames( 'node') # "SUID" "shared name"
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
